-
Notifications
You must be signed in to change notification settings - Fork 0
CCI RL interaction
simoncmo edited this page Jul 2, 2022
·
2 revisions
- 7/2/2022
# 1. Cell trek but remove cell that generated from Celltrek itself
celltrek_obj_snonly = RemoveCelltrekDuplication(celltrek_obj)
# 2. make graph
graph_snonly = MakeCellTrekDelaunayGraph(celltrek_obj_snonly, max_distance = 500)
# 3. Make Plot object
sample_use = 'HT2610C1'
cell_type_plt = 'Tumor'
km_use = 3
palette_use_cluster = MakeClusterPalette(k = km_use, celltype = cell_type_plt)
palette_use = c('') # No palette
cell_column = 'cell_type_RNA'
plot_obj_snonly = PrepareSpatialPlotObject(celltrek_obj_snonly,
graph_snonly,
km_use = 3,
palette_cluster = palette_use_cluster,
cell_column = cell_column,
palette = palette_use, celltype_highlight = cell_type_plt, n_order = 1)
# 4. Plots - Basic plots
p_ct_full_network = MakeDelaunayGraphPlot(plot_obj_snonly, show_all_points = T)
p_ct_neighbor_network = MakeDelaunayGraphPlot(plot_obj_snonly, show_neighbor_pts = F, show_target_pts = T) %>%
AddClusterHighlight(plot_obj_snonly)
p_ct_neighbor_profile = MakeNeighborProfilePlotV2(plot_info_obj = plot_obj_snonly)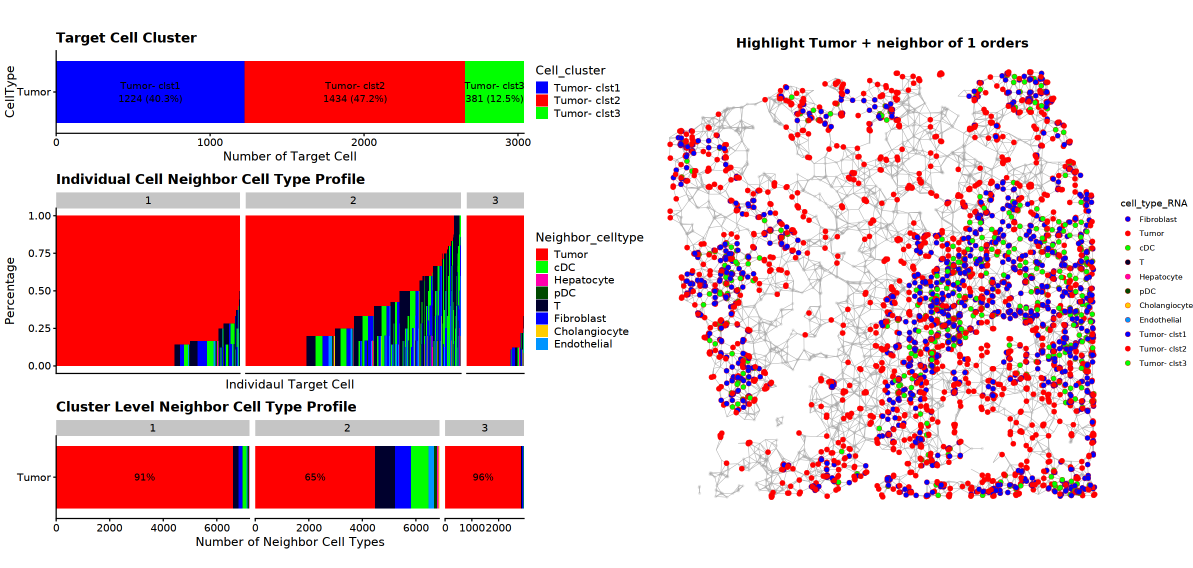
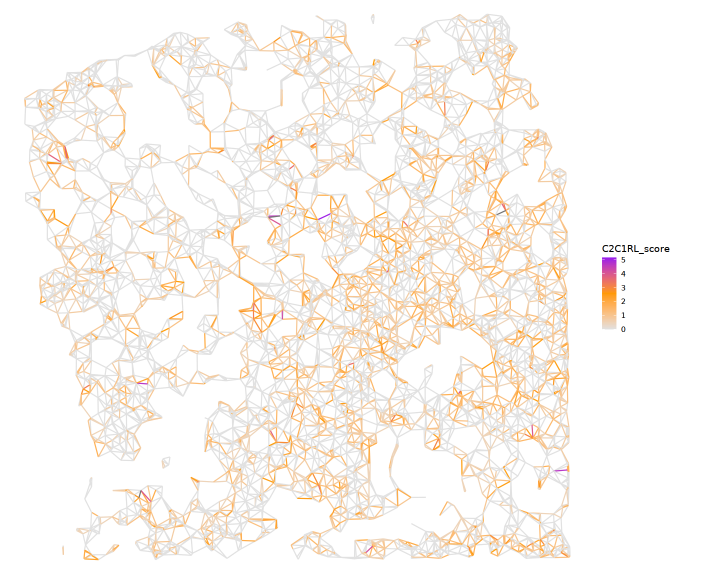
Note: R-L score is the multiplication of Cell1-Receptor expression * Cell2-Ligand expression
(Or could be Cell2-Receptor expression * Cell1-Ligand expression depends on direction parameter)
# 0a. preprocess
## sn keep only cell in CellTrek
sn_only = RemoveScRNANotInCellTrek(scRNA_obj = sn, CellTrek_obj = celltrek_obj)
## Add SCT score to cell trek obj
## Need this for RL analysis
celltrek_obj_snonly = AddAssayToCellTrek(celltrek_obj = celltrek_obj_snonly, scRNA = sn_only, assay = 'SCT')
# 0b. Get edge info
# Add Edge related info
plot_obj_snonly = GetEdgeCellTypeTable(plot_obj_snonly) %>% AddEdgeGroup
## 1. Get RL scores
# 2. pairs as example
r_gene = 'LAMB1'
l_gene = 'ITGB1'
plot_obj_snonly = GetRLscores(celltrek_obj_snonly,
plot_obj_snonly,
r_gene, l_gene)
plot_obj_snonly = GetRLscores(celltrek_obj_snonly, plot_obj_snonly,
'CCR4', 'CCL4')
## Plot RL score
MakeCellTrekRLscore(plot_obj_snonly, 'LAMB1_ITGB1', direction = 'C2C1')